Twisting and turning to target antibiotic resistance
Research Press Release | June 16, 2016
Researchers are getting closer to understanding how some natural antibiotics work so they can develop drugs that mimic them.
A recent review commissioned by the British government reported that, “without policies to stop the worrying spread of antimicrobial resistance, today’s already large 700,000 deaths every year would become an extremely disturbing 10 million every year by 2050, more people than currently die from cancer.”
Some natural antibacterial products are known to target an enzyme, called MraY, which is essential for the synthesis of an important component of the bacterial cell wall. Despite the promise of this approach, researchers have not been able to develop synthetic antibacterial products that target this enzyme, largely because they do not have a clear understanding of its structure.
A team of scientists from Hokkaido University in Japan and Duke University in the U.S. have found that MraY undergoes remarkable changes in its structure to bind to a natural inhibitor called MD2.
Normally, MraY interacts with a substance called UM5A in the bacterial cell wall, ultimately producing peptidoglycan, which gives the cell wall its structural strength. The research team found that MraY interacts with the natural inhibitor MD2 in a way that is different from its interaction with UM5A.
The team crystallized MraY while bound to MD2 in order to determine its structure and how the two molecules interact. They also investigated the results of making genetic changes to MD2’s structure and how this affected its interaction with MraY.
Previously, it was thought that MD2 interacted with MraY by mimicking the action of UM5A. This study, published in the journal Nature, showed that a part of MraY unwound and rearranged itself to create a pocket that allowed a part of MD2, which has no equivalent in UM5A, to bind to the enzyme like a two-pronged plug inserting into a socket. The study also showed that another part of the MD2 structure further anchors it to MraY, increasing the inhibitor’s affinity to the enzyme.
“The large conformational arrangement observed in MraY indicates conformational plasticity,” the researchers write. This could be the reason why MraY accommodates so many structurally different inhibitors, they say.
These findings help add to the knowledge and understanding necessary to develop new drugs that target bacteria.
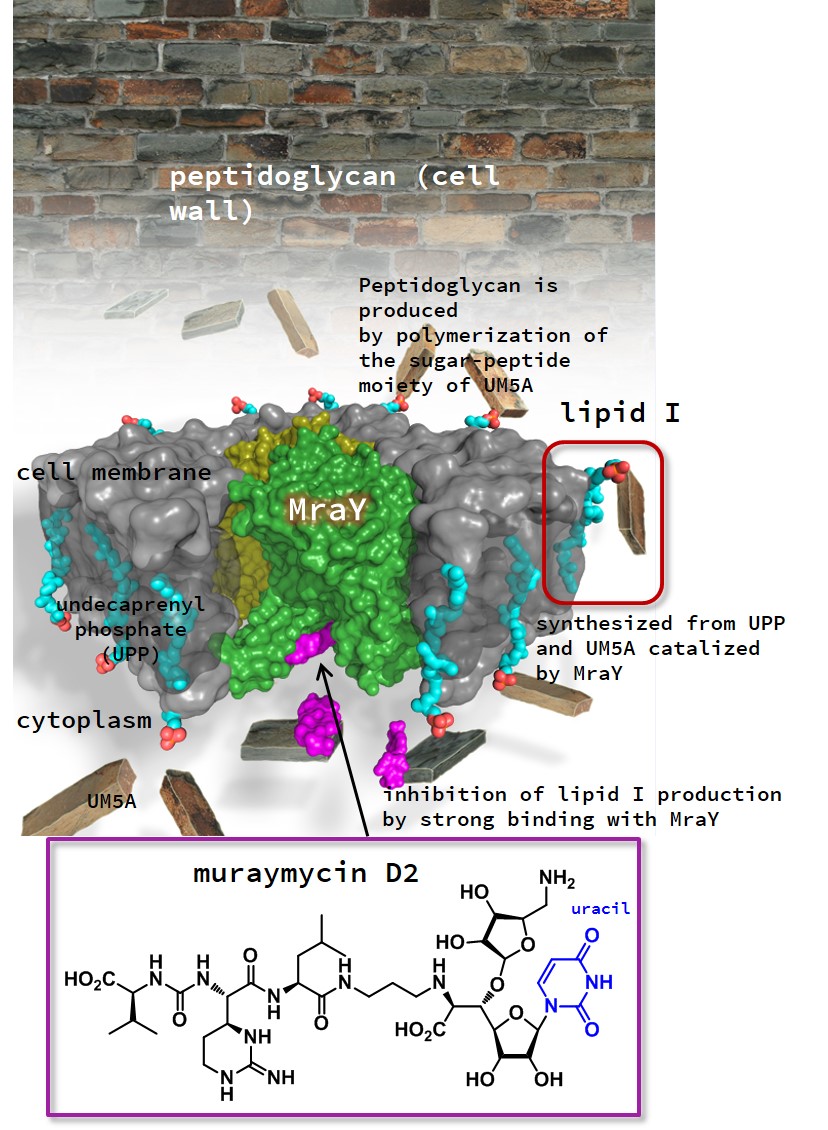
Structure of the complex of muraymycin D2 and MraY,
an enzyme responsible for biosynthesis of peptidoglycan
MD2 (purple) binds to MraY (green) like a two-pronged plug inserting into a socket. This ultimately inhibits the formation of a substance that gives the bacterial cell wall its structural strength.
Original article
Structural insights into inhibition of lipid I production in bacterial cell wall synthesis, Nature (4.18.2016)
Contacts:
Professor Satoshi ICHIKAWA
Center for Research and Education on Drug Discovery
Hokkaido University
Email: ichikawa[at]pharm.hokudai.ac.jp
Mr. Naoki NAMBA (Media Officer)
Global Relations Office
Office of International Affairs
Hokkaido University
Email: pr[at]oia.hokudai.ac.jp
Tel: +81-11-706-8034
