New biosynthetic pathway unique to pathogenic microorganisms
Research Press Release | July 07, 2017
Scientists have discovered a unique biosynthetic pathway for D-Glutamate, an important cell wall component in pathogenic bacteria, which could lead to the development of medicines and agricultural chemicals.
Cell walls (peptideglycan) are essential for microorganisms to maintain cell integrity. The team led by Professor Toru Dairi, from Hokkaido University’s Graduate School of Engineering, focused on the biosynthesis of D-Glutamate (Glu), which is a component of cell walls. In general, D-Glu is supplied by an enzyme called Glu racemases. But the team discovered through its comparative genomics of various microorganisms that Xanthomonas oryzae—a pathogenic bacteria that causes rice plant leaves to turn white before killing the plant—does not have Glu racemase genes.
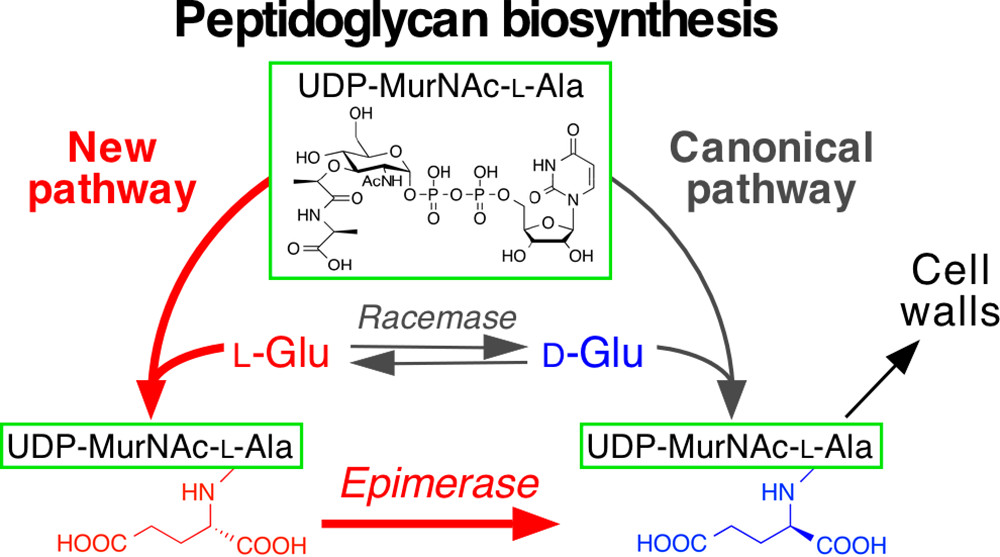
To figure out what is compensating for the missing piece, the researchers conducted shotgun cloning experiments in which the genes of X. oryzae were introduced to Escherichia coli mutants that cannot produce D-Glu. The team found out the two genes created a pathway that bypassed the defect, enabling the production of D-Glu. In general, D-Glu binds with a molecular complex called UDP-MurNAc-L-Ala, but in this pathogenic microorganism, L-Glu bound with the complex mediated by one of the two genes they identified. The other gene then epimerizes (isomerizes) L-Glu to D-Glu.
“Like X.oryzae, other pathogenic bacteria such as Stenotrophomonas which trigger in-hospital infections could also take the newly discovered pathway. We hope that the pathway will provide new targets for developing medicines and agricultural chemicals against bacteria,” says Toru Dairi.
Original article:
Dairi T. et al., A glycopeptidylidyI-glutamate epimerase for bacterial peptidoglycan biosynthesis, Journal of the American Chemical Society, March 15, 2017.
DOI: 10.1021/jacs.7b01221
Contacts:
Professor Toru Dairi
Laboratory of Applied BioChemistry
Graduate School of Engineering
Hokkaido University
Email: dairi[at]eng.hokudai.ac.jp
Naoki Namba (Media Officer)
Global Relations Office
Institute for International Collaboration
Hokkaido University
Tel: +81-11-706-2185
Email: pr[at]oia.hokudai.ac.jp