Working in the wild to help protect animals and humans from zoonoses
Research Highlight | January 21, 2021
This article is an excerpt from the research magazine “Tackling Global Issues vol.3 Fighting the menace of zoonosis.” Click here to see the table of contents.

Hirofumi Sawa, MD, Ph.D.
Professor and Deputy Director of the Research Center for Zoonosis Control (CZC)
Research Area: Molecular pathobiology, virology, epidemiology, animal model
Hirofumi Sawa started his career three decades ago working as a clinical cardiologist conducting basic research at Hokkaido University’s Graduate School of Medicine. He soon acquired a keen interest in viruses, leading him to start research on JC polyomavirus, which causes fatal demyelinating disease of the central nervous system, under Dr. Kazuo Nagashima. Therefore, in 2005, it was a natural progression for him to apply for a professorship at the university’s Research Center for Zoonosis Control (CZC).
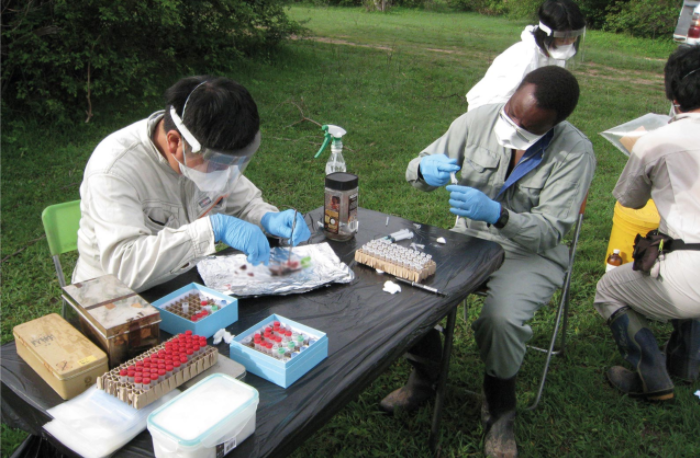
Sawa leaves his CZC lab three to four times a year to conduct research in the field in Zambia, catching mosquitos and collecting wildlife and athropods in caves. Indeed, his work style fluctuates from being a researcher at state-of-the-art medical institutions at Hokkaido University and Washington University in St. Louis to working in the wild where there is no electricity or other modern comforts.
“I have always navigated my career path to a direction that has interested me,” Sawa said. “I aspired to be a medical doctor because I wanted to cure patients. Now I am devising preemptive measures against zoonoses. Broadly speaking, this has the same goal. My strategy is just a bit different from many of my colleagues.”
Discovering new zoonosis viruses
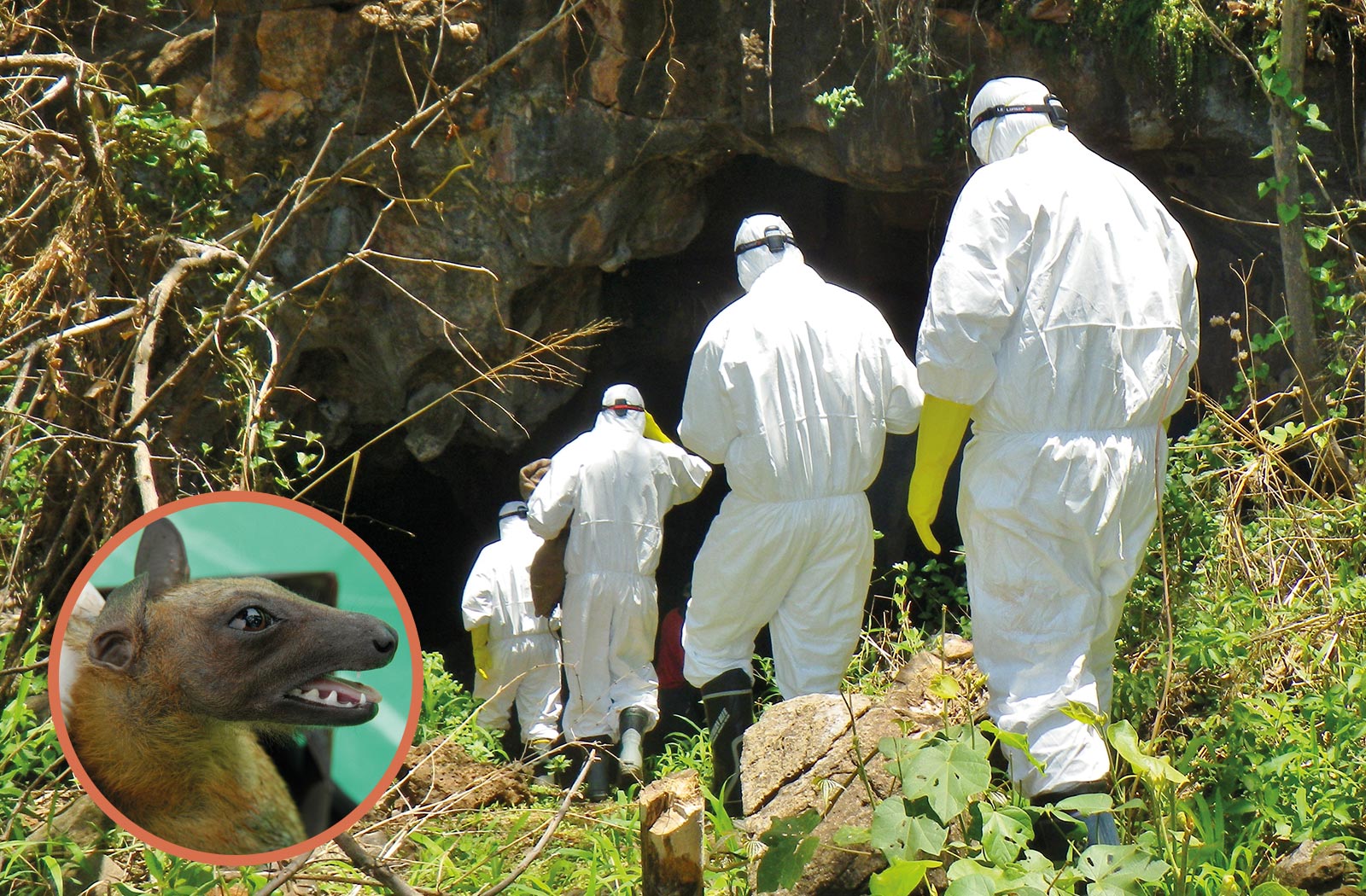

Sawa and his colleagues enter a cave in Zambia to collect bat feces. The inset shows a fruit bat (Eidolon helvum).
Sawa’s research base in Zambia is the Hokudai Center for Zoonosis Control in Zambia (HUCZCZ), which was established in 2007 by Hokkaido University’s CZC as a hub for epidemiological studies in sub-Saharan Africa. HUCZCZ is located at the University of Zambia’s School of Veterinary Medicine.
Pathogens that pose a huge threat to public health, such as the human immunodeficiency virus (HIV), Ebola virus and Zika virus, are believed to have originated in Africa, where yellow fever and malaria are also endemic. This makes Africa a key continent for conducting research on infectious diseases.
“If we want to control zoonoses, it is important to conduct research in developing countries where outbreaks of known and unknown infectious diseases are reported,” Sawa said. “It is difficult to eradicate infectious diseases, but we can prevent zoonoses from spreading across the globe if we implement evidence-based preventive measures.”
To that end, Sawa said, “In addition to unravelling the diversity of viruses carried by animals, we also need to search for new viruses, and determine their host range, pathogenicity and possible transmission routes. This task deserves proper attention considering that over 70 percent of emerging infectious diseases originated from wildlife.”
Sawa’s research team has been credited with detecting and isolating more than 20 new viruses carried by wild animals and arthropods.
For example, Sawa’s colleague Dr. Michihito Sasaki, who is a lecturer at Hokkaido University, detected a virus belonging to the paramyxoviridae family from feces of wild baboons in Zambia. Other members of this family include two well-known viruses — measles virus and mumps virus, which are mainly associated with human respiratory disorders. Detection of paramyxoviruses in wild baboons was made possible by employing a method called reverse transcriptase-polymerase chain reaction (RT-PCR) with the use of degenerate primers, which is simply a mixture of primers that are similar, but not identical. Using this method, his research team identified one of the viruses as human parainfluenza type 3 (HPIV3), which is most closely related to HPIV3 strain Riyadh 149/2009, isolated from a hospitalized child in Saudi Arabia in 2009.
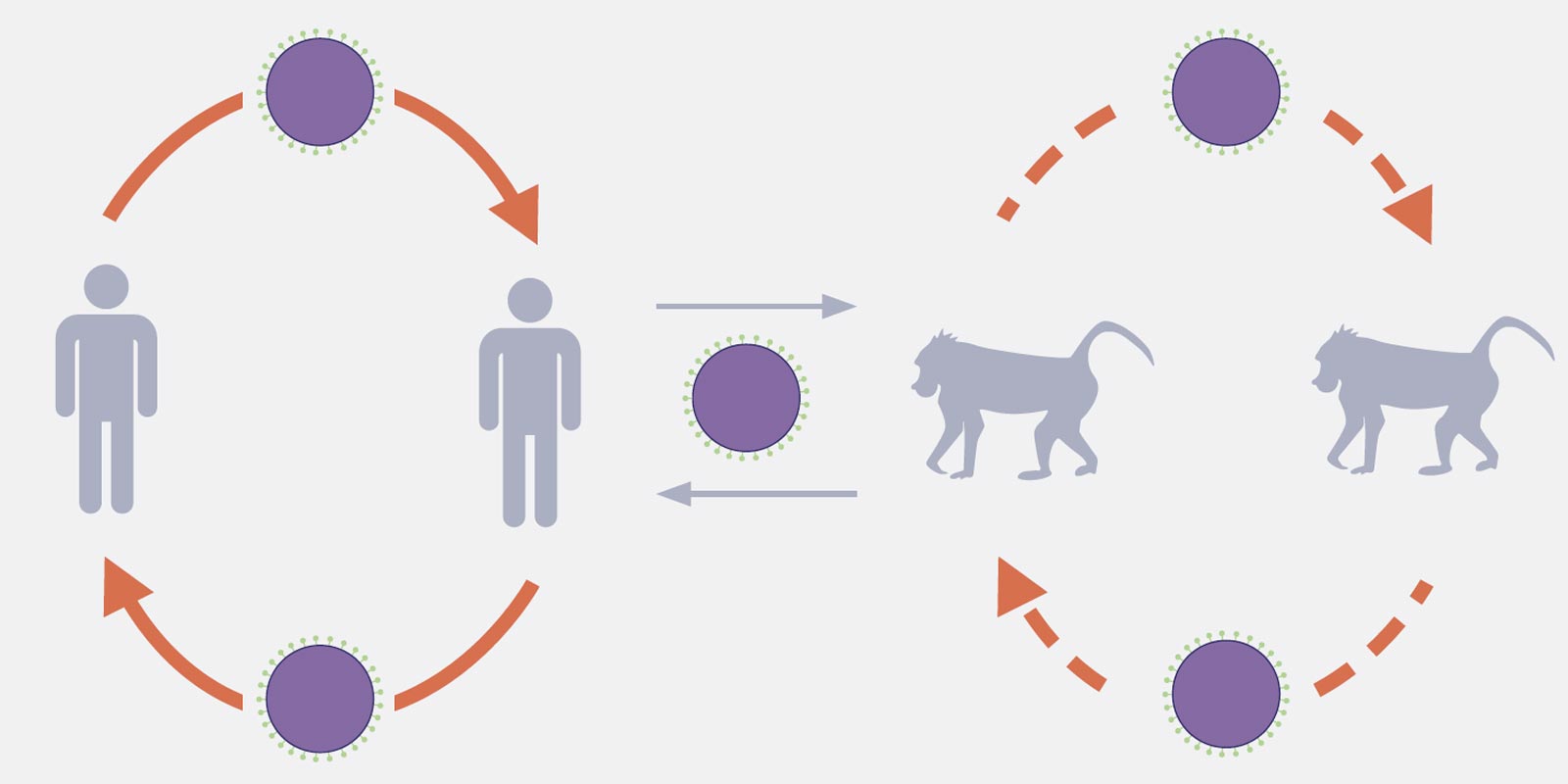
“We believe that the virus is transmitted from baboons to humans and from humans to other animals,” Sawa said. “Baboons tend to come close to humans in search of food. Because of this close contact, the virus can be transmitted from baboons to humans and vice versa.”
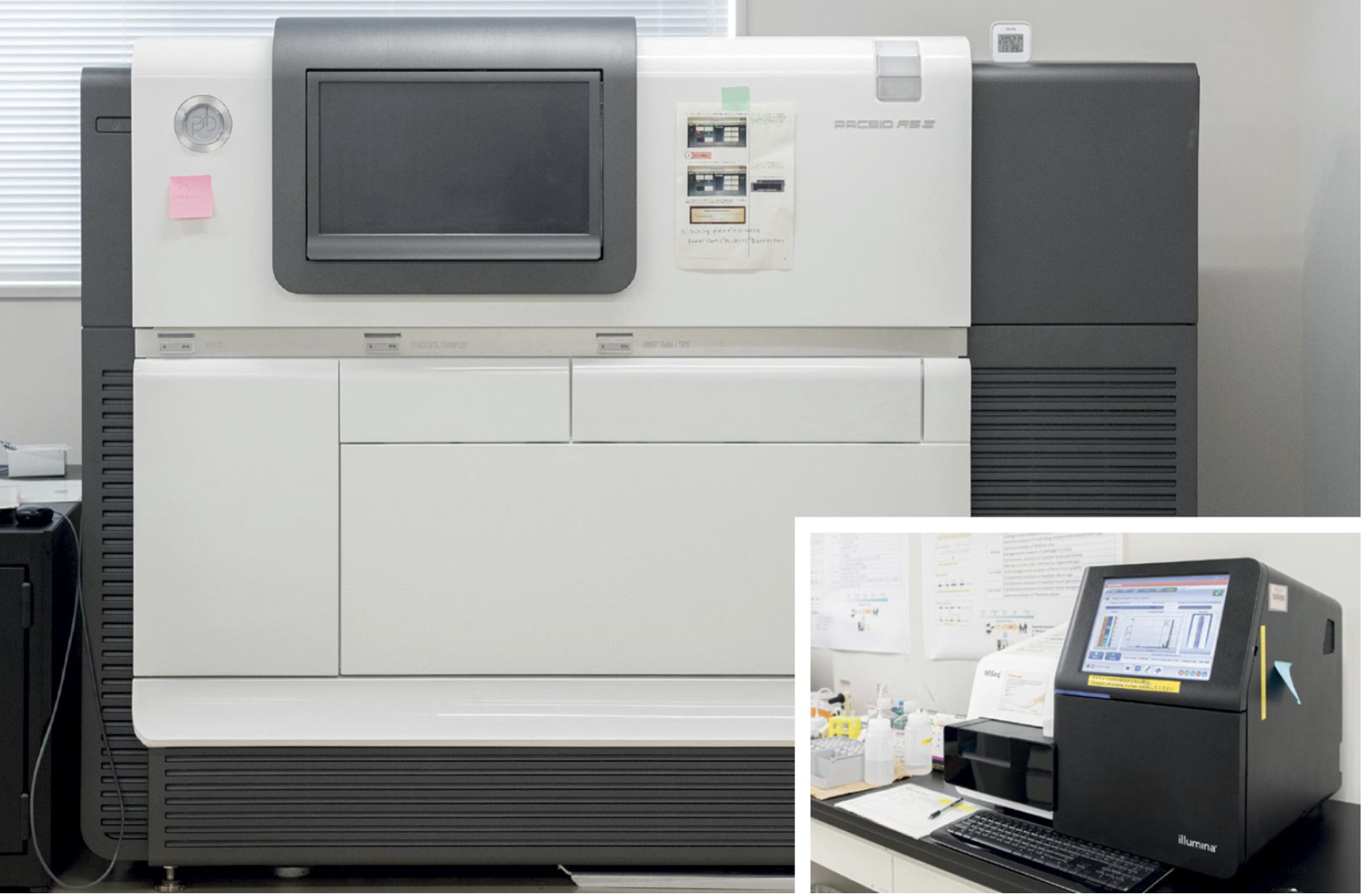
Next-generation sequencers for comprehensive genomic profiling
According to Sawa, next-generation sequencing (NGS), which enables sequencing of DNA and RNA much more quickly and comprehensively than previous techniques, has revolutionized genomic and molecular biology. RT-PCR (used for analyzing RNA viruses) and PCR (used for analyzing DNA viruses) are instrumental when the targeted virus is known, but NGS becomes a powerful tool when researchers do not know exactly what kind of virus they are dealing with. NGS allows for a comprehensive and cost-effective sequencing of known and unknown pathogens, such as DNA and RNA viruses.
Thanks to these new technologies, the number of virus species acknowledged by the International Committee on Taxonomy of Viruses increased from 1,898 in 2005 to 6,590 as of May 2020.
Specifically, Sasaki employs shotgun metagenomics, an approach that involves shearing DNA or RNA extracted from samples and subsequent sequencing of the small fragments using NGS. The generated information is analyzed using bioinformatics tools. Now, the team has a convenient, portable sequencing device capable of detecting viruses in the field.
Using the latest technologies, the team detected Group A rotavirus in insect-eating bat species from Zambian caves. Rotaviruses have a double-stranded RNA genome consisting of eleven segments. Rotaviruses detected by Sasaki had six genome segments highly similar (96 percent to 99 percent) to the human rotaviruses, while the remaining five segments were similar to those of rotaviruses found in livestock and wild animals. “The result suggests that rotaviruses carried by bats have a similar genetic profile to those found in humans. This is also evidence of cross-species transmission and probable genetic reassortment being behind the high genetic diversity seen in rotaviruses,” said Sawa.
Furthermore, Sasaki detected several new viruses in insect-eating white-toothed shrews, which inhabit some rural residential areas in Zambia. A new picornavirus found in shrews was similar to human picornaviruses, which cause gastroenteritis. Additionally, his team detected a new cyclovirus that was markedly different from other known cycloviruses found in animal feces. This new cyclovirus was very similar to viruses detected in the cerebrospinal fluid of encephalomyelitis patients.
“These findings suggest that white-toothed shrews, which are found widely around the globe, carry viruses unique to them, and can, therefore, be a potential host to zoonotic viruses,” Sawa warned.
“Collecting samples from wildlife and isolating viruses from them are time-consuming tasks, but they are integral parts in the fight against zoonoses,” Sawa said. To shorten the process, Sawa and his collaborators are developing a system to effectively isolate viruses by making optimal cells needed for replication of viruses and then rotary-culturing cells and viruses together to boost viral replication.
Mosquitos the “most dangerous insect”
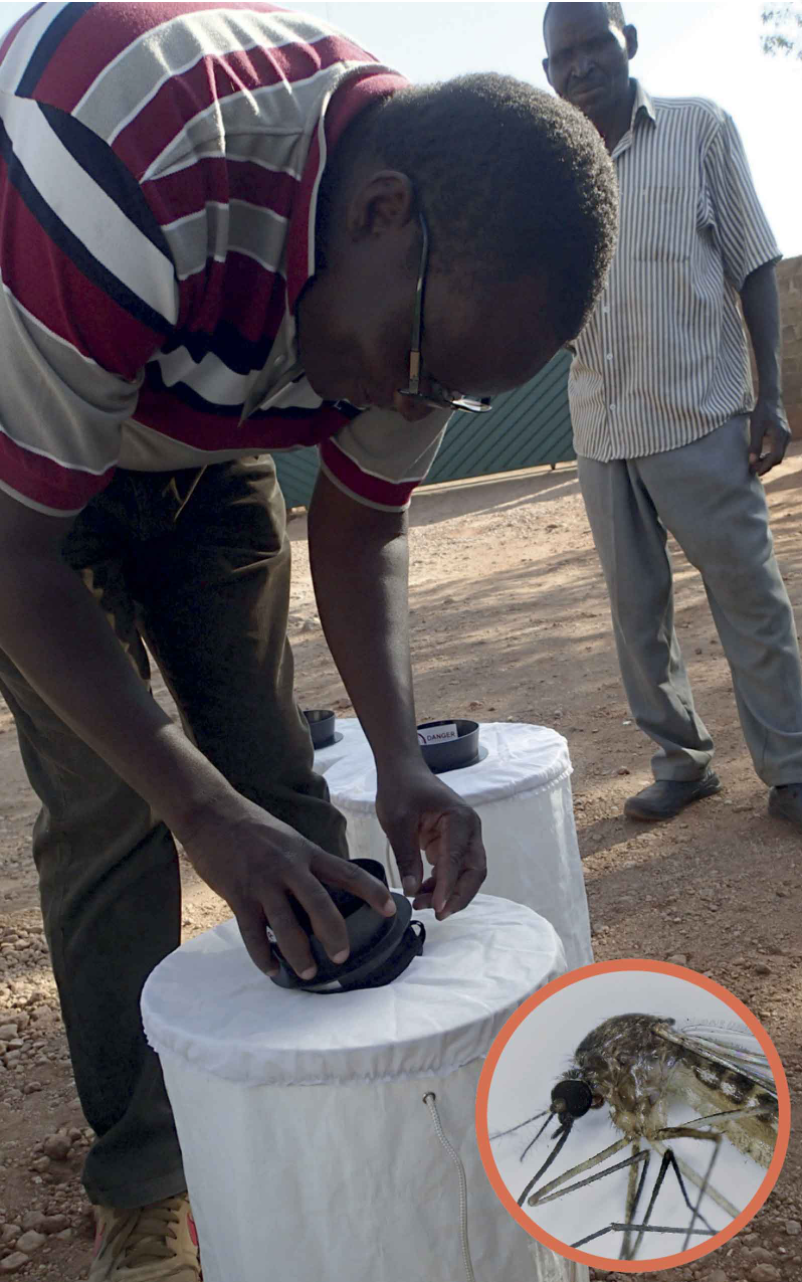

A total of 17,826 female mosquitos were gathered between
2012 and 2019 in Zambia. Traps that produce odor attractant and a common mosquito Culex quinquefasciatus (inset) that was found to be a vector of West Nile virus.
Dr. Yasuko Orba, who works with Sawa, focuses on virus surveillance in mosquitos, which are widely considered to be the world’s most dangerous insect because they are a vector for many diseases. From 2012 to 2019, Orba collected more than 17,800 female mosquitos from different places in Zambia in search of known and unknown viruses. This effort culminated in the first isolation of West Nile virus in Zambia. Orba also surveyed birds, horses and humans based on the West Nile virus transmission route:
from mosquitos to birds; birds to mosquitos; and finally to horses and humans.
“Preemptive measures can be taken based on research on the diversity of viruses wild animals and arthropods carry, as well as studies on the natural hosts and pathogenicity of viruses,” Sawa said. For instance, Sawa is helping one of his colleagues, Dr. Akihiko Sato from Shionogi & Co., Ltd., to develop an antiviral drug to treat arthropod-borne viruses (arboviruses) and coronaviruses.
“As a doctor conducting basic research on human cardiovascular diseases, I never imagined that I would visit Zambia to research infectious diseases,” Sawa said. “It’s not easy, but I’ve been able to keep up because I like Zambian people for their kindness and humility.”
Click here to see the table of contents.