Mice possess natural gene therapy system
A previously mysterious small RNA molecule in mice is found to play a crucial role in gene expression, and may be the first identified member of a new class of regulatory RNAs.
RNA (ribonucleic acid) is best known as the messenger RNA (mRNA) that carries a copy of a gene’s information out from the cell nucleus to where it can be decoded to make protein molecules. But RNA also performs other key functions, including the regulation of gene activity by a variety of small non-coding RNAs—those whose genetic sequence is not used to generate proteins.
One such non-coding RNA is the small RNA called 4.5SH, found only in small rodents including mice and rats. It is produced from multiple copies of its gene, leading to the accumulation of up to 10,000 copies of the RNA molecule per cell.
A team of researchers led by Professor Shinichi Nakagawa at Hokkaido University has discovered a new role for 4.5 SH RNA—circumventing mutations in mouse DNA during the maturation of mRNA. Their findings were published in the journal Molecular Cell.
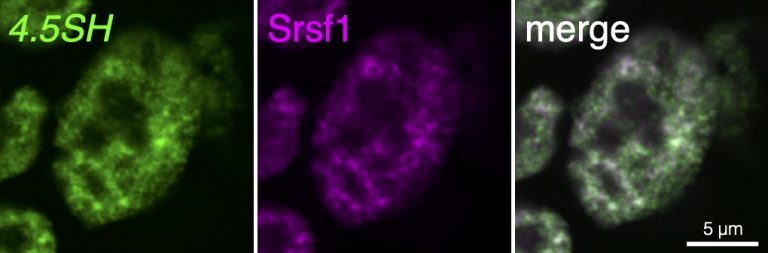
“4.5SH RNA was discovered in the 1970s, yet despite its abundance and presence in many types of tissues, its function had remained a mystery for over 40 years,” says Nakagawa.
To understand its role, the researchers created mutations in mouse embryos that abolished 4.5SH production, discovering that this caused early death at the embryo stage.
“It was known that the mouse genome has many lethal mutations in genes that code essential proteins,” explains Nakagawa. “4.5 SH RNA has the ability to detoxify these mutations in bulk—essentially, it is a natural gene therapy to protect against mutations.”
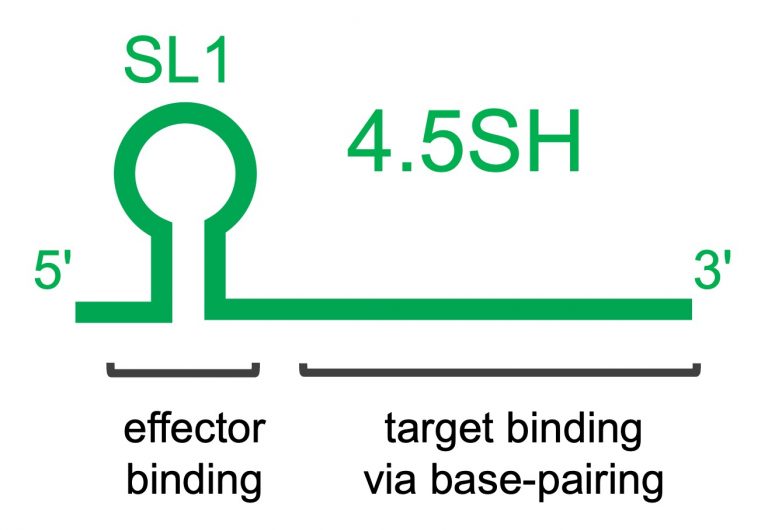
Analysis of the structure of 4.5SH RNA showed that it is composed of two modules. One serves as a sensor to find abnormal sequences, and the other brings in a tool that prevents the incorporation of the abnormal sequences into mRNA by a process called alternative splicing.
“To our knowledge, this is the first example of a naturally produced RNA that can regulate alternative splicing in a definitive on/off manner,” says Nakagawa. “Our research also suggests that a substantial portion of such non-coding RNAs may be involved in controlling alternative splicing.”
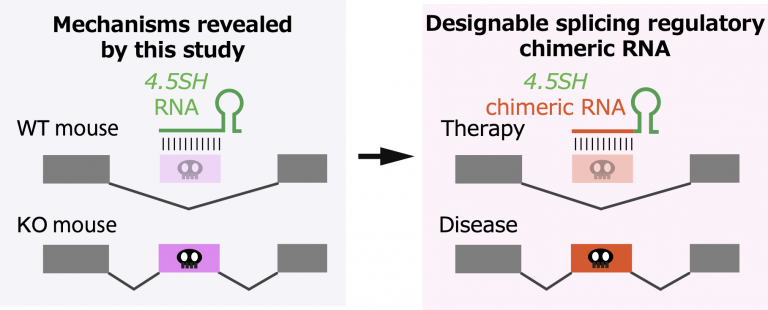
The researchers were also able to use 4.5SH to design a programmable molecular system that could manipulate splicing in cells in selected ways. This might become a new and useful tool for genetic engineering.
“Our discovery suggests the possibility of developing new gene therapy drugs that recognize only specific genetic mutations by modifying the sensor module of 4.5 SH RNA, so we may be able to prevent toxic regions associated with disease from being expressed,” Nakagawa explains.
Original Article:
Rei Yoshimoto, et al. 4.5SH RNA counteracts deleterious exonization of SINE B1 in mice. Molecular Cell. December 13, 2023.
DOI: 10.1016/j.molcel.2023.11.019
Funding:
This work was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI (JP16H06276, 21H05274, 21K19246, 22K05565, 21H00253, 22H02545, JP20H05784); the Japan Agency for Medical Research and Development (AMED; JP23ym0126801); and the Global Facility Center (GFC), Hokkaido University.
Contacts:
Professor Shinichi Nakagawa
Faculty of Pharmaceutical Sciences
Hokkaido University
Tel: +81-11-706-3745
Email: nakagawas[at]pharm.hokudai.ac.jp
Sohail Keegan Pinto (International Public Relations Specialist)
Public Relations & Communications Division
Office of Public Relations and Social Collaboration
Hokkaido University
Tel: +81-11-706-2186
Email: en-press[at]general.hokudai.ac.jp
